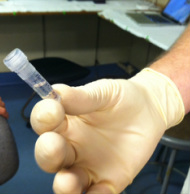
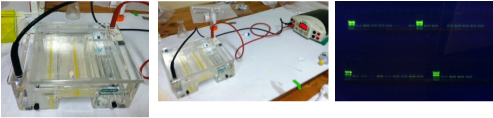
 Gold! Gold! Hard to believe but this tiny bottle contains genetic information that will allow us to characterize the diversity of the microbial community at Station ALOHA. This was the final product from 5 days of sampling and many hours filtering, extracting, priming, amplifying and pipetting! We took water samples from the surface of the water all the way through the water column. We are interested in determining the diversity of microbial organisms at Station Aloha. We used PCR (polymerase chain reaction) to amplify pieces of DNA across several orders of magnitude. Each sample received a specific primer which serves as "tag" so we can link the microbial community to a depth and time of day they were collected. Primers are short DNA fragments that contain sequences complementary to the target region along the DNA polymerase. We checked to see whether the PCR generated the anticipated DNA fragments (sometimes called amplicons) using a process called agarose gel electrophoresis, which separates the PCR products based on size. A DNA ladder containing DNA fragments of known sizes is run alongside the PCR products. Once we determined that the PCR worked for each sample we then conducted a procedure that pooled all of the samples together and we ended up with one small bottle. It is truly mind blowing to consider how much data and information is contained in such a small volume!
11 Comments
9/3/2013 08:32:40 pm
Message in a bottle...I am so grateful to find your particular post. I have bookmarked this website and I will keep visiting you for further such interesting posts.keep on posting...
Reply
really awesomr post11 waititng for more updates
Reply
11/6/2020 03:59:38 am
I appreciate what you folks are up as well. This sort of shrewd work and inclusion! Keep up the extraordinary works folks I've added you folks to my own blogroll.
Reply
10/6/2021 09:34:01 am
A great website with interesting and unique material what else would you need.
Reply
2/22/2022 01:45:11 am
I trust every one of you are having an amazing week's end. I added another overview. This one is more humble, but simultaneously important. I think the accompanying one will be more noteworthy.
Reply
11/22/2022 01:24:12 am
I really like to read your blog very much.
Reply
bulksmscoimbatore
12/8/2022 02:45:14 am
Reply
12/22/2023 10:05:24 am
Hi my friend! I want to say that this article is amazing, great written and include almost all vital infos. I would like to see extra posts like this.
Reply
12/26/2023 05:51:26 am
I appreciate, cause I found exactly what I was looking for. You’ve ended my four day long hunt! God Bless you man. Have a great day.
Reply
Leave a Reply. |
Sarah Q. FosterSarah is a 2nd year Ph.D. student in the Fulweiler Lab. This blog documents her experience taking a summer course "Microbial Oceanography: From genome to biome" at C-MORE at the University of Hawai'i at Manoa. ArchivesCategories |

 RSS Feed
RSS Feed